COEVOL Multi-Scale Coevolution
Living systems are highly integrated, with a multitude of levels of organization, from molecular and intra-cellular scales to ecosystems. Complex organisms are themselves consortia of macro- and micro-organisms, which work together with their host to build the individual. Yet, each of these organisms can function and evolve in the short term according to its own logic, possibly in conflict with other higher or lower levels, or with other time scales. The once common idea among evolutionists that natural selection results in organisms perfectly adapted to their environment is now severely undermined. Not only because, as the Red Queen explains to Alice, one has to run relentlessly to keep its place in a changing environment, or because past evolutionary history and chance constrain the possibilities of present adaptation, but also because different levels of selection have interests that are generally difficult to reconcile.
Multi-scale coevolution resets classical questions in evolutionary biology
One example, of particular interest is the question of the source of heritable variations. The phenotype of organisms in a population is influenced not only by variations in their nuclear and mitochondrial genomes, the dynamics of which is the object of population genetics, but also more and more patently by the consortium of microbes and genetic elements that constitute its microbiome and virome. The hologenome designates this complex assembly of genetic materials, which obey different rules of transmission and different evolutionary strategies. The ability of symbionts to manipulate host phenotypes or to interfere with each other influences the evolutionary dynamics of all players in ways that are yet poorly understood. In addition, new questions arise, such as the importance of co-adaptation in these systems and their consequences in maintaining cohesive biological systems.
- Symbiosis: a response to and a source of divergent selection
Using a variety of approaches combining experimental evolution, genomic, functional, phenotypic and behavioral data, we aim to test whether symbiosis facilitates diversification and to characterize the underlying microevolutionary processes.
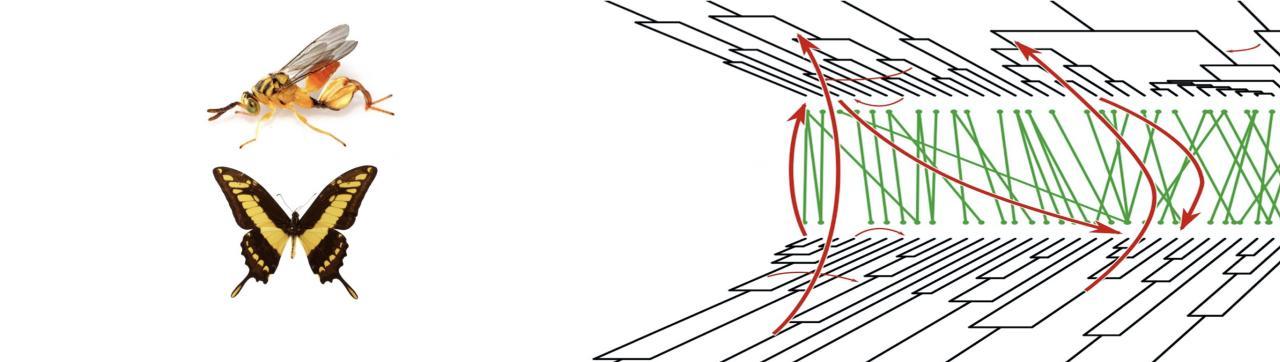
- Ecological networks of horizontal gene transfer
We develop original methods to detect gene transfer and we investigate the factors that influence the routes of gene transfers among microbes but also among insects.
- The interplay between symbiosis, infection and immunity and its evolutionary consequences
We try to understand the intimate interaction of hosts with pathogens, symbionts and transposable elements and how it affects the extended phenotype of the host.
- Transgenerational inheritance and environment changes
We try to decipher the molecular mechanisms that underlie rapid adaptation to environment and to test for transgenerational inheritance of fitness traits.
- Intragenomic conflicts and demography
We are developing models to test whether changes in the demography of the host affect the dynamics of transposable elements.
- The determinism of phenotypic convergence
We study the genomic basis of convergent phenotypic evolution in particular in the case of animals and plants adaptation to increasing temperature and decreasing water.
- Reconciling the tree of life
We develop phylogenetic methods for “reconciling” gene/species or host/symbiont histories and use these methods to explore the bulk of extinct or undescribed species and the history of association of symbiotic microbes with their hosts.
Integrating methods
The methods we use to tackle the questions raised by multi-scale co-evolution extend from theory, modelling and simulation to big data analysis, lab (notably on insects), and to a lesser extent, field activities.
Implication of research, responsibility of researchers and citizen sciences
From our research (some of which have immediate consequences in health, agriculture and ecology) and our concerns about the responsibility of scientists in society, we are committed to promote an “implicative” research. The implicative position means that we try to work on the link between science and society, not only through a one-way communication, applying or explaining our science, but also favoring early discussions on research projects, that may influence our research directions.
Publications
Display of 661 to 690 publications on 710 in total
Phenotypic plasticity of sternopleural bristle number in temperate and tropical populations of Drosophila melanogaster
Genetical Research . 81 : 25-32
Journal article
see the publicationStrain-specific regulation of intracellular Wolbachia density in multiply infected insects
Molecular Ecology . 12 : 3459-3465
Journal article
see the publicationG+C3 structuring along the genome: a common feature in prokaryotes
Molecular Biology and Evolution . 20 : 471-483
Journal article
see the publicationFrom gene trees to organismal phylogeny in prokaryotes: the case of the gamma-Proteobacteria
PLoS Biology . 1 ( 1 ) : 101-109
Journal article
see the publicationThe source of laterally transferred genes in bacterial genomes
Genome Biology . 4 : R57.1-R57.12
Journal article
see the publicationVariation of the genome size estimate with environmental conditions in Drosophila melanogaster
Cytometry - Part A . 55A : 43-49
Journal article
see the publicationInfectious behavior in a parasitoid
Science . 302 ( 5652 ) : 1930
Journal article
see the publicationBetween- and within-host species selection on cytoplasmic incompatibility-inducing Wolbachia in haplodiploids
Evolution - International Journal of Organic Evolution . 57 : 421-427
Journal article
see the publicationPhylogenetics and the cohesion of bacterial genomes
Science . 301 ( 5634 ) : 829-832
Journal article
see the publicationOn a modular domination game
Theoretical Computer Science . 306 : 291-303
Journal article
see the publicationWorldwide distribution of transposable element copy number in natural populations of Drosophila simulans
Evolution - International Journal of Organic Evolution . 57 : 159-167
Journal article
see the publicationWORLDWIDE DISTRIBUTION OF TRANSPOSABLE ELEMENT COPY NUMBER IN NATURAL POPULATIONS OF DROSOPHILA SIMULANS
Evolution - International Journal of Organic Evolution . 57 ( 1 ) : 159-167
Journal article
see the publicationStimulating effects of the insecticide chlorpyrifos on host searching and infestation efficacy of a parasitoid wasp.
Pest Management Science . 58 ( 4 ) : 321-328
DOI: 10.1002/ps.454
Journal article
see the publicationStimulating effects of the insecticide chlorpyrifos on host searching and infestation efficacy of a parasitoid wasp
Pest Management Science . 58 : 321-328
Journal article
see the publicationInfection polymorphism and cytoplasmic incompatibility in Hymenoptera-Wolbachia associations
Heredity . 88 : 361-365
Journal article
see the publicationPhylogeny of six African Leptopilina species (Hymenoptera : Cynipoidea Figitidae) parasitoids of Drosophila spp. with descriptions of three new species
Annales de la Société Entomologique de France . 38 : 319-332
Journal article
see the publicationA phylogenomic approach to bacterial phylogeny: evidence of a core of genes sharing a common history
Genome Research . 12 : 1080-1090
DOI: 10.1101/gr.187002
Journal article
see the publicationEvolution of genome size in Drosophila. is the invader`s genome being invaded by transposable elements?
Molecular Biology and Evolution . 19 : 1154-1161
Journal article
see the publicationPartial compensation of the sublethal effect of deltamethrin on the sex pheromonal communication of Trichogramma brassicae
Chemosphere . 42 : 985-991
Journal article
see the publicationModification by the Insecticide Chlorpyrifos of the Behavioral Response to Kairomones of a Parasitoid Wasp Leptopilina boulardi.
Archives of Environmental Contamination and Toxicology . 41 : 436-442
Journal article
see the publicationRemoving symbiotic Wolbachia bacteria specifically inhibits oogenesis in a parasitic wasp
Proceedings of the National Academy of Sciences of the United States of America . 98 : 6247-6252
Journal article
see the publicationWithin-species diversity of Wolbachia-induced cytoplasmic incompatibility in haplodiploid insects
Evolution - International Journal of Organic Evolution . 55 : 1710-1714
Journal article
see the publicationTransposable elements and genome evolution of invasive species: the case of Drosophila
Genetics Selection Evolution . 33 : S107-S120
Journal article
see the publicationRemoving the symbiotic bacteria Wolbachia inhibits oogenesis in a parasitic wasp.
Proceedings of the National Academy of Sciences of the United States of America . ( 98 ) : 6247-6252
Journal article
see the publicationDivergence between Drosophila santomea and allopatric or sympatric populations of D. yakuba using paralogous amylase genes and migration scenarios along the Cameroon volcanic line
Molecular Ecology . 10 : 649-660
Journal article
see the publicationConnected Joins in Graphs
International Conference on Integer Programming and Combinatorial Optimization (IPCO) . 2081 : 383-395
Conference paper
see the publicationEvidence for female mortality in Wolbachia-mediated cytoplasmic incompatibility in haplodiploid insects: Epidemiologic and evolutionary consequences
Evolution - International Journal of Organic Evolution . 54 : 191-200
Journal article
see the publicationLes Wolbachia bactéries endosymbiotiques parasites de la reproduction des Arthropodes : Circulation diversité et évolution des effets dans un complexe parasitaire
incollection . -- : 95-99
Journal article
see the publicationGenotype-environment interaction for quantitative trait loci affecting life span in Drosophila melanogaster
Genetics . 154 : 213-227
Journal article
see the publication